Project webpage: https://auditore.cab.inta-csic.es/solid/
Publications: https://auditore.cab.inta-csic.es/solid/publications/
Principal investigators: Víctor Parro García IP1 and Mercedes Moreno Paz IP2
Current team: Víctor Parro García (CAB), Mercedes Moreno Paz (CAB), Alberto González Fairén (CAB), Laura Sánchez García (CAB), David Ruano Gallego (CAB), María Ángeles Lezcano Vega (CAB), Rita Dos Santos Severino (CAB), Pedro Mustieles del Ser (Univ. Complutense de Madrid / CAB), Miriam García Villadangos (CAB), Andoni Moral Inza (Flight Segment Dept., INTA), Juan Manuel Manchado Ortega (CAB), Raquel López Heredero (Space Optics Dept., INTA), José Antonio Rodríguez-Manfredi (CAB), Javier Gómez-Elvira (Payloads Dept., INTA)
Project of the Biomolecules in planetary exploration group
Since its inception, the Centro de Astrobiología has been working on the design and construction of SOLID (Signs Of LIfe Detector), an instrument based on an antibody microarray, created for the detection and identification of microorganisms and biochemical compounds by in situ analysis of solid residues (soil, ground rock or ice) and liquid samples.
SOLID is the materialisation of the project “Detection of Biomolecules in Planetary Exploration”, which has been continuously funded by the Spanish Ministry of Science and Innovation since 2004. SOLID’s working hypothesis is based on the assumption that, under similar environmental conditions, organisms respond and adapt with similar molecular strategies and mechanisms. If terrestrial environments analogous to other planetary bodies are known and understood and some of the life forms and debris they leave behind can be characterised, and systems for detecting them developed, then this methodology could be extrapolated to the search for life in planetary exploration.
Several prototypes have been developed and we now have the SOLID3.1 version, capable of detecting microbial markers and large molecular weight compounds (mainly biological polymers such as proteins, polysaccharides, nucleic acids) as well as low molecular weight compounds. The core of SOLID is an antibody microarray biosensor, called LDChip (Life Detector Chip), capable of hosting more than 300 different antibodies to detect a similar number of potential biomarkers. We currently have more than 450 antibodies against bacterial and archaeal strains, natural polymeric crude extracts, proteins, polysaccharides, nucleic acids, and small molecules such as amino acids, or polycyclic aromatic hydrocarbons.
SOLID consists of two functional and physically separate units in version 3.1: the Sample Preparation Unit (SPU) for sample reception, homogenisation, processing, and filtering; and the Sample Analysis Unit (SAU), which houses the LDChip (Life Detector Chip) antibody biosensor microarray for immunological assays. The current instrument design consists of a SPU3.0 for 10 samples, or a SPU3.1 of a 10-fold reusable extraction cell. The SAU module contains the fluidics, optics, and 5 LDChip replicates for as many analyses. Considering this block diagram, the mass distribution is: SPU3.0 about 5.5 kg, SPU3.1 4.5 kg, and SAU about 2 kg.
The current proposal “Detecting Signals of Life in Planetary Exploration (SOLID)” (2019-2021) is based on three main pillars: (1) Advancing the science supporting the SOLID concept for the search for complex organic molecules (biotic and abiotic) as potential targets (aromatic amino acids, prebiotic peptides, other molecules) to be searched for in planetary exploration; (2) The scientific-technological challenge of implementing the LDChip biosensor with the selection of the best performing antibodies from our collection and the production of new ones, as well as continuing the tests already initiated in space-relevant conditions and their validation in campaigns in terrestrial environments; (3) Technological activities to improve the performance of the different components and subsystems of the current version of the instrument (SOLID3. 1) and its testing in space-relevant conditions in order to increase the TRL (Technological Readiness Level) and to be able to successfully opt for future proposals to search for life in planetary exploration.
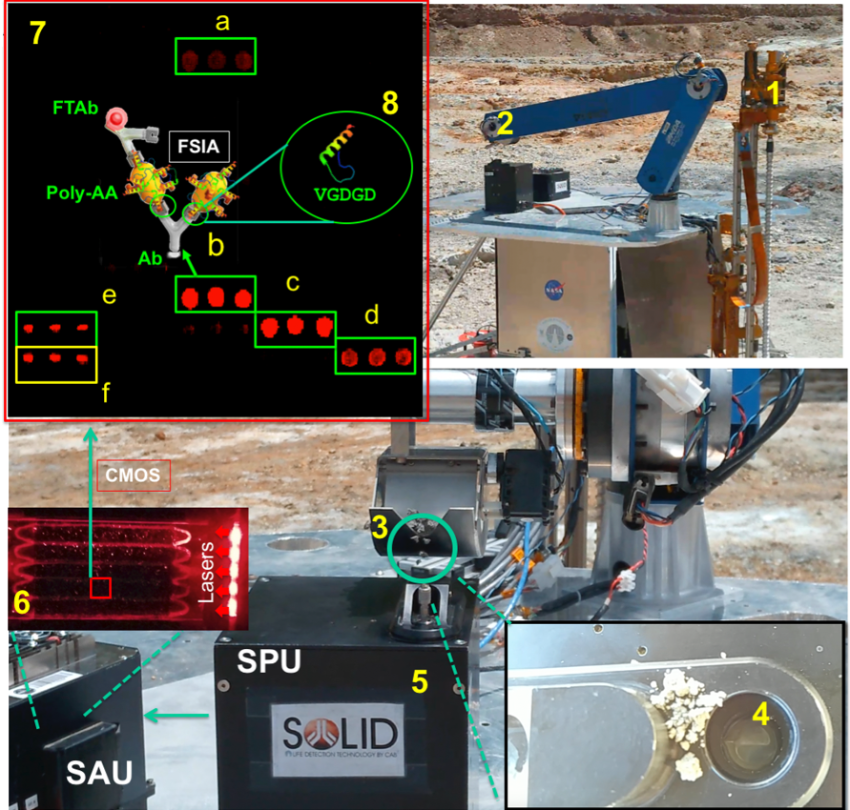
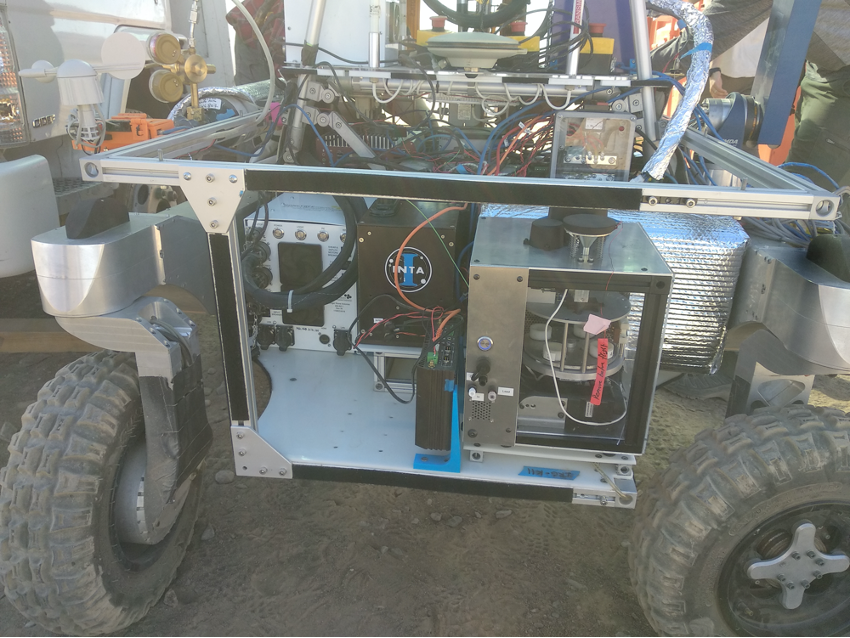
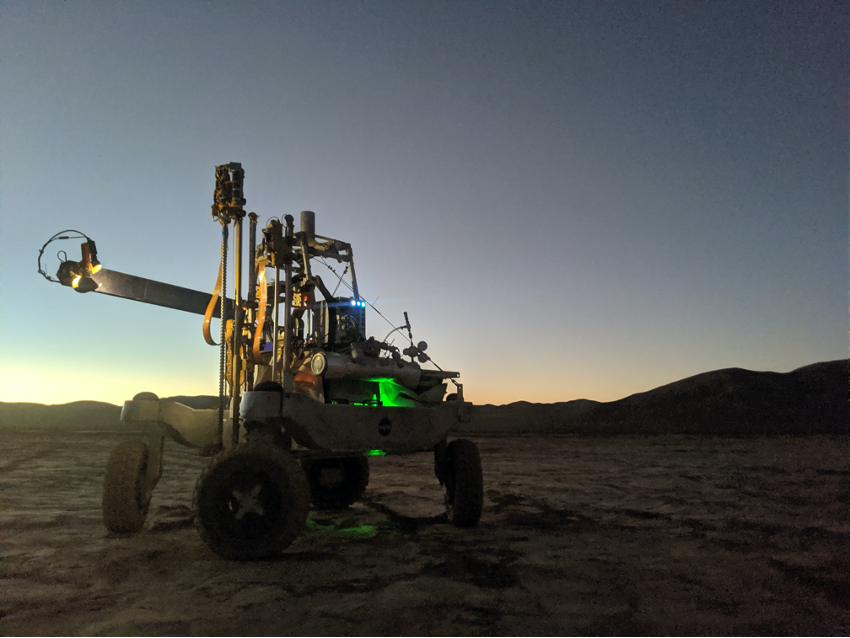
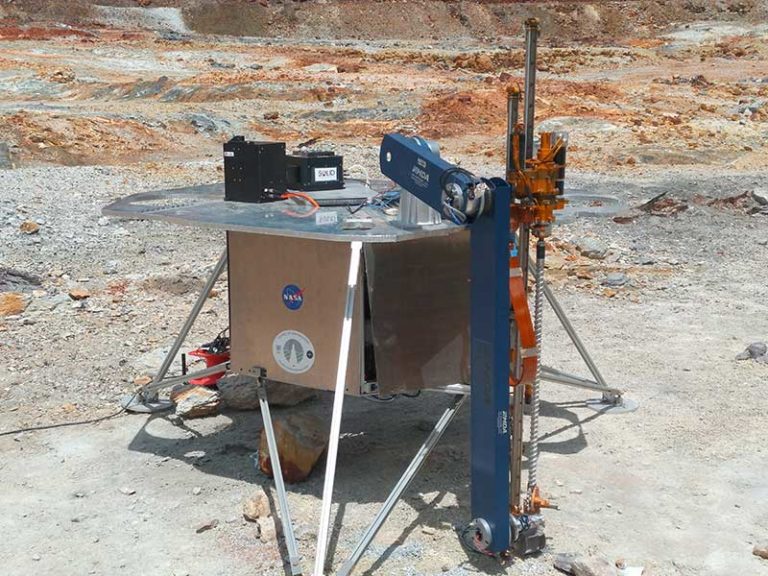
This is how SOLID works: (1, 2) SOLID on the IceBreaker simulated platform during the NASA-Ames collaboration campaign at Rio Tinto in 2017. The IceBreaker(1) drill extracts the subsurface sample (up to 1 m deep) and the robotic arm (2) feeds SOLID (3) into the extraction cell of the SPU. At this point the remote and automated operation is activated: the extraction solution (4) is added, closed and homogenised/extracted by ultrasonication (5), after 10 micron filtration it is incubated with LDChip in the SPU (6). After development with fluorescent antibodies, an image is obtained with a CCD camera (6), downloaded and analysed on “Earth” (7). Fluorescent three-point replicas (a-e) reveal the presence of microbial biomarkers, many of them protein fragments (peptides) as shown in the scheme (8). Ab, capture antibody immobilised on LDChip; FTAb, fluorescent tracer antibody; FSIA, Fluorescent Sandwich ImmunoAssay. Each LDChip spot is 120-150 microns in diameter and contains approximately 10 million antibody molecules waiting to capture molecular structures for which they were “trained”. These are polyclonal antibodies, to increase the likelihood of capturing biomarkers.
Campaigns
Both LDChip and SOLID have been tested in the laboratory and in multiple field campaigns in some of the best and most representative terrestrial analogues of Mars, such as Antarctica, Rio Tinto (Huelva), the Atacama Desert (Chile), or the Chilean Andes. During the collaborative campaign with NASA-Ames Research Center ARADS2019 in the Atacama Desert, a fully remote SOLID operation simulation was carried out, with the operations centre in Madrid and San Francisco. SOLID-LDChip images were received via satellite and the science team analysed them and decided on the procedure to follow for new samples.